Experimental Design
Knowing the hypothesis testing framework is necessary but not sufficient. An experiment that's correctly analyzed but poorly designed will give you narrow confidence intervals around the wrong thing, or require three months of runtime to detect a lift you expected to see in two weeks. Experimental design is where most of the leverage lives: choosing the right randomization unit (user vs. session vs. city), applying variance reduction techniques that can halve your required sample size, constructing ratio metrics carefully enough that your test is valid, and selecting metrics that are sensitive to the changes you care about. A well-designed experiment is one where the randomization is clean, the metric responds quickly to the change under test, and the variance is as low as possible given the data already collected before the experiment started.
Theory
CUPED subtracts the pre-experiment covariate component θX̃ from Y, shrinking variance without changing the expected treatment effect.
Before you run a single experiment, you can cut the required sample size in half — without collecting any new data. Pre-experiment measurements of the same metric already contain most of the user-level variance; subtracting the predictable component leaves only the noise the treatment actually needs to overcome. CUPED does exactly this: last week's behavior controls for this week's baseline, shrinking distributions without touching the causal estimate.
Randomization as identification
The magic of a randomized experiment is that random assignment makes the treated and control groups statistically identical in expectation — not just on observed covariates, but on every characteristic including unobserved ones. This is what gives A/B tests their causal identification power.
Formally: under random assignment, , so:
No other assumption required. This is why experiments are the gold standard and why observational methods need untestable assumptions to claim causal effects.
CUPED variance reduction
The key lever for running shorter experiments is reducing variance without changing sample size. Controlled-experiment Using Pre-Experiment Data (CUPED) (Deng et al., Microsoft, 2013) achieves this by subtracting the component of predictable from a pre-experiment covariate :
The formula is the OLS slope from regressing on — the unique coefficient that minimizes over all possible linear adjustments. Any other value of gives an unbiased treatment effect estimate but higher variance; this is the variance-minimizing choice. The unbiasedness holds regardless of , which is why CUPED can be applied post-hoc at readout time rather than pre-specified before the experiment.
The variance of the adjusted outcome is:
where . For Daily Active Users (DAU) with week-over-week stability :
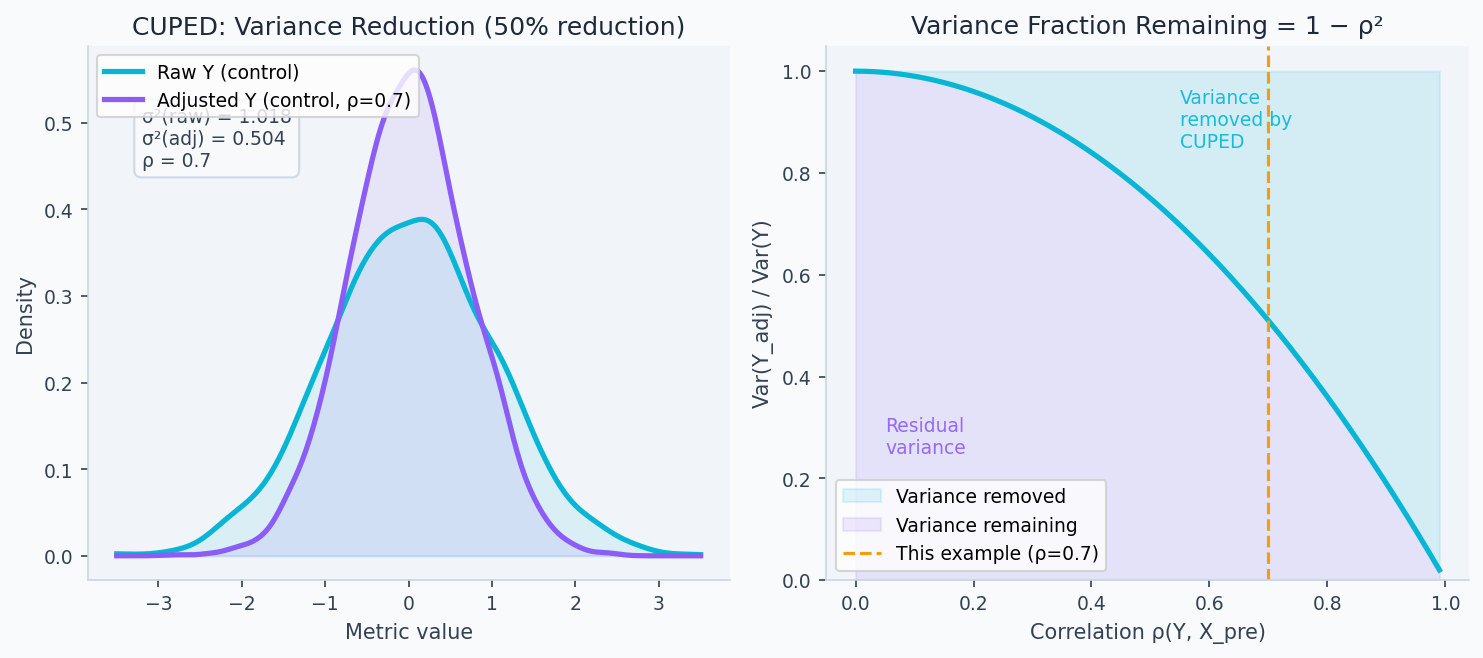
Same required number of standard errors, 64% less variance → 64% fewer users needed → 64% shorter experiment. The left panel below shows exactly this: the adjusted distribution is narrower, concentrating more probability mass near the true mean. The right panel shows that the variance remaining grows rapidly as falls — if your pre-experiment covariate has no predictive power, CUPED doesn't help.
Key properties:
- Estimate on the pooled dataset (both arms together), never per-arm — this avoids post-treatment contamination
- The adjustment is unbiased regardless of : for any . Optimizing only minimizes variance.
- Can be applied post-hoc at readout time if you forgot to pre-apply it
Ratio metrics and the delta method
Business metrics are often ratios: revenue per session = total revenue / total sessions. The naive variance formula ignores the denominator's variability. The correct formula via the delta method:
The covariance term matters. If sessions increase alongside revenue (treatment increases both numerator and denominator), the naive estimator will overstate variance. If sessions decrease while revenue increases, it will understate variance.
Design effect for cluster randomization
When SUTVA is violated — a treatment affecting one user spills over to others — you randomize at a higher level: cities, devices, households. The design effect (DEFF) quantifies the cost of cluster randomization versus individual randomization:
where is average cluster size and is the intraclass correlation coefficient (ICC). For city-level experiments with users/city and :
You need approximately twice as many users compared to user-level randomization. The effective sample size is . The ICC is often underestimated — cluster experiments are chronically underpowered as a result.
Stratified randomization
Rather than pure random assignment, stratified randomization assigns units within pre-specified strata (e.g., new vs. returning users, mobile vs. desktop). The variance reduction:
The reduction equals the between-stratum variance in outcomes divided by . For a metric where new vs. returning users differ significantly (large ), stratification gives meaningful variance reduction with no statistical cost.
Walkthrough
CUPED implementation
import numpy as np
import pandas as pd
from scipy import stats
def apply_cuped(
df: pd.DataFrame,
outcome_col: str,
pre_experiment_col: str,
variant_col: str = 'variant',
) -> pd.Series:
"""
CUPED-adjusted outcome.
theta estimated on the full dataset (not per-arm) to avoid post-treatment bias.
"""
X = df[pre_experiment_col].values
Y = df[outcome_col].values
# Estimate theta = Cov(Y, X) / Var(X) on pooled data
theta = np.cov(Y, X)[0, 1] / np.var(X)
# Adjust
Y_adj = Y - theta * (X - X.mean())
return pd.Series(Y_adj, index=df.index, name=f"{outcome_col}_cuped")
def variance_reduction_from_cuped(Y: np.ndarray, X: np.ndarray) -> dict:
"""How much variance does CUPED remove for this metric/covariate pair?"""
rho = np.corrcoef(Y, X)[0, 1]
return {
"rho": round(rho, 3),
"variance_reduction_pct": round((1 - rho**2) * 100, 1),
"sample_size_reduction_pct": round((1 - rho**2) * 100, 1), # same factor
}
# Example: pre-experiment revenue as covariate for post-experiment revenue
df['revenue_cuped'] = apply_cuped(df, 'revenue_post', 'revenue_pre')
stats_pre = variance_reduction_from_cuped(df['revenue_post'].values, df['revenue_pre'].values)
# rho=0.72 → 48% variance reduction → 48% fewer users neededSample size with adjustments
import math
def sample_size_suite(
baseline_rate: float,
mde_relative: float,
rho_cuped: float = 0.0,
avg_cluster_size: float = 1.0,
icc: float = 0.0,
alpha: float = 0.05,
power: float = 0.80,
) -> dict:
"""
Sample size accounting for CUPED variance reduction and cluster design effect.
"""
z_alpha = stats.norm.ppf(1 - alpha / 2)
z_beta = stats.norm.ppf(power)
delta = baseline_rate * mde_relative
p_pool = baseline_rate + delta / 2
sigma = math.sqrt(2 * p_pool * (1 - p_pool))
n_base = math.ceil(((z_alpha + z_beta) * sigma / delta) ** 2)
# Apply CUPED variance reduction
n_cuped = math.ceil(n_base * (1 - rho_cuped**2))
# Apply design effect for clustering
deff = 1 + (avg_cluster_size - 1) * icc
n_clustered = math.ceil(n_cuped * deff)
n_clusters = math.ceil(n_clustered / avg_cluster_size) if avg_cluster_size > 1 else None
return {
"n_unadjusted": n_base,
"n_with_cuped": n_cuped,
"n_with_clustering": n_clustered,
"n_clusters_needed": n_clusters,
"design_effect": round(deff, 2),
"days_needed": math.ceil(n_clustered / (200_000 * 0.5)), # example traffic
}
# Pure individual randomization: 45% baseline, 2% MDE
r = sample_size_suite(0.45, 0.02)
print(f"No adjustments: {r['n_unadjusted']:,} per arm") # 19,652
# With CUPED (rho=0.7)
r = sample_size_suite(0.45, 0.02, rho_cuped=0.7)
print(f"With CUPED: {r['n_with_cuped']:,} per arm") # 10,718 (45% fewer)
# City-level randomization, 50 users/city, ICC=0.02
r = sample_size_suite(0.45, 0.02, avg_cluster_size=50, icc=0.02)
print(f"Cluster: {r['n_clusters_needed']:,} clusters needed") # 390Ratio metric variance (delta method)
def ratio_metric_stats(
numerator_A: np.ndarray, # e.g., revenue per user in control
denominator_A: np.ndarray, # e.g., sessions per user in control
numerator_B: np.ndarray,
denominator_B: np.ndarray,
) -> dict:
"""
Compare ratio metrics (e.g., revenue per session) between arms.
Uses delta method for variance of ratio.
"""
def ratio_stats(num, den):
r = num.mean() / den.mean()
n = len(num)
var_ratio = (1/den.mean()**2) * (
np.var(num)
- 2 * r * np.cov(num, den)[0,1]
+ r**2 * np.var(den)
) / n
return r, var_ratio
r_A, var_A = ratio_stats(numerator_A, denominator_A)
r_B, var_B = ratio_stats(numerator_B, denominator_B)
delta = r_B - r_A
se = math.sqrt(var_A + var_B)
z = delta / se
p_value = 2 * (1 - stats.norm.cdf(abs(z)))
ci = (delta - 1.96 * se, delta + 1.96 * se)
return {
"ratio_control": round(r_A, 4),
"ratio_treatment": round(r_B, 4),
"delta": round(delta, 4),
"relative_lift_pct": round(delta / r_A * 100, 2),
"ci_95": (round(ci[0], 4), round(ci[1], 4)),
"p_value": round(p_value, 5),
}Randomization unit choice
The unit you randomize on determines what questions you can answer:
| Unit | When to use | Tradeoffs |
|---|---|---|
| User ID | Most experiments — measures per-user effects | Cannot control SUTVA violations between users |
| Session ID | When user-level effects aren't relevant (e.g., anonymous search) | Same user may see both arms — "dilution" |
| Device ID | When the product is device-specific | Multi-device users dilute effects |
| City / Market | When SUTVA is violated (marketplace, social) | Cluster design effect reduces power significantly |
| Time period | Switchback experiments for marketplace pricing | Carry-over effects between periods |
Always hash the randomization unit deterministically:
import hashlib
def assign_variant(unit_id: str, experiment_id: str, split: float = 0.5) -> str:
"""Deterministic, consistent assignment. Same user always gets same variant."""
key = f"{experiment_id}:{unit_id}".encode()
bucket = int(hashlib.sha256(key).hexdigest(), 16) % 10_000 / 10_000
return "treatment" if bucket < split else "control"Using a hash keyed on both experiment_id and unit_id ensures independence across experiments — the same user isn't always in the treatment arm.
Pre-experiment AA test
Simulate 1000 AA tests (no actual treatment) on historical data. Under a correctly specified variance model, exactly ~5% should yield p < 0.05. A higher false positive rate reveals a broken variance model.
def run_aa_simulation(
user_metrics: np.ndarray,
n_simulations: int = 1000,
alpha: float = 0.05,
) -> dict:
"""
Validate variance model by simulating AA tests on historical data.
"""
false_positives = 0
for _ in range(n_simulations):
idx = np.random.permutation(len(user_metrics))
mid = len(idx) // 2
_, p = stats.ttest_ind(user_metrics[idx[:mid]], user_metrics[idx[mid:]])
if p < alpha:
false_positives += 1
fpr = false_positives / n_simulations
return {
"false_positive_rate": round(fpr, 3),
"expected": alpha,
"verdict": "PASS" if abs(fpr - alpha) < 0.02 else "FAIL — check variance model",
"likely_cause": (
"clustering effects or autocorrelation" if fpr > alpha * 1.5 else "ok"
),
}Analysis & Evaluation
Where Your Intuition Breaks
Running a longer experiment always fixes an underpowered one. Sample size is determined by variance, not time. If your metric is dominated by high-variance users (e.g., a small fraction of users drive most revenue), doubling experiment duration barely helps — each new day adds the same noisy users. The fix is variance reduction (CUPED, winsorization, switching to a lower-CV metric), not more time. Duration matters only to the extent that it determines how many independent users you observe; for low-traffic products, even a month of runtime may fall short of the required n.
Metric selection: sensitivity vs. relevance
Not all metrics are equally useful as primary metrics. Two dimensions:
Sensitivity (statistical): how much signal per user. Measured by coefficient of variation (). Lower CV → smaller sample size for the same MDE.
Relevance (business): how predictive is this metric of long-term business outcomes? A metric can be easy to move (sensitive) but irrelevant (click-through on an interstitial you're about to remove).
The right primary metric maximizes both. Estimate relevance empirically: compute the correlation between candidate short-term metrics and long-term outcomes on historical data. Compute sensitivity using pre-experiment variance.
| Metric | CV (lower is better) | Corr with 6-month retention |
|---|---|---|
| Sessions/week | 0.8 | 0.71 |
| Revenue/session | 2.1 | 0.54 |
| Search success rate | 0.4 | 0.83 ← good primary |
| Feature adoption | 0.6 | 0.31 |
Variance estimation from historical data
Use 4 weeks of pre-experiment data to estimate . Apply a variance inflation factor of 1.1–1.2 to be conservative — this accounts for:
- Users not yet in the experiment behaving differently from those who are
- Week-of-month effects not captured in 4 weeks
- Cross-device users counted once but generating multiple events
For highly right-skewed metrics (revenue, session duration), winsorize at the 99th percentile before estimating variance. The same winsorization threshold must be applied at analysis time.
Production-Ready Code
In production, CUPED runs as a post-processing step on the raw event data after experiment close — it doesn't require changing instrumentation. The key engineering decision is covariate selection: compute correlations on pre-experiment data and pick the highest-rho columns automatically. Assignment must use a deterministic hash so the same user always gets the same variant across sessions, devices, and service calls.
# production_experiment_design.py
# CUPED pipeline with automated covariate selection, sample size service,
# and deterministic variant assignment.
from __future__ import annotations
import hashlib, math
from typing import Sequence
import numpy as np
import pandas as pd
import scipy.stats as stats
# ── Covariate selection ───────────────────────────────────────────────────────
def select_cuped_covariates(
df: pd.DataFrame,
outcome_col: str,
candidate_cols: Sequence[str],
min_rho: float = 0.3,
) -> list[str]:
"""
Returns pre-experiment columns with |rho| >= min_rho, sorted by descending |rho|.
Only pass columns measured BEFORE experiment start as candidates.
Typical candidates: last-week metric, last-month metric, days-since-signup.
"""
Y = df[outcome_col].values
ranked = []
for col in candidate_cols:
rho = float(np.corrcoef(Y, df[col].values)[0, 1])
if abs(rho) >= min_rho:
ranked.append((col, rho))
ranked.sort(key=lambda x: abs(x[1]), reverse=True)
return [col for col, _ in ranked]
# ── Multi-covariate CUPED via OLS residualisation ────────────────────────────
def apply_cuped_pipeline(
df: pd.DataFrame,
outcome_col: str,
covariate_cols: Sequence[str],
variant_col: str = "variant",
) -> pd.Series:
"""
Generalises single-covariate CUPED to multiple covariates via OLS.
theta is estimated on the POOLED dataset — never per arm — to avoid
post-treatment contamination.
Returns the adjusted outcome series indexed like df.
"""
Y = df[outcome_col].values
X = df[covariate_cols].values
X_dm = X - X.mean(axis=0) # demean each covariate
theta, *_ = np.linalg.lstsq(X_dm, Y, rcond=None) # OLS minimises Var(Y_adj)
Y_adj = Y - X_dm @ theta
return pd.Series(Y_adj, index=df.index, name=f"{outcome_col}_cuped")
def cuped_variance_reduction(
df: pd.DataFrame,
outcome_col: str,
covariate_cols: Sequence[str],
) -> dict:
"""How much variance does the covariate set explain? Use to justify covariate choice."""
Y = df[outcome_col].values
X = df[covariate_cols].values
X_dm = X - X.mean(axis=0)
theta, *_ = np.linalg.lstsq(X_dm, Y, rcond=None)
Y_adj = Y - X_dm @ theta
reduction = 1 - Y_adj.var() / Y.var()
return {
"covariates": list(covariate_cols),
"variance_reduction_pct": round(reduction * 100, 1),
"sample_size_reduction_pct": round(reduction * 100, 1),
}
# ── Sample size service ───────────────────────────────────────────────────────
def sample_size_service(
baseline: float,
mde_relative: float,
rho_cuped: float = 0.0,
avg_cluster_size: float = 1.0,
icc: float = 0.0,
alpha: float = 0.05,
power: float = 0.80,
) -> dict:
"""
Single entry point for all experiment sizing. Accounts for CUPED variance
reduction and cluster design effect in sequence.
"""
z_a = stats.norm.ppf(1 - alpha / 2)
z_b = stats.norm.ppf(power)
delta = baseline * mde_relative
p_pool = baseline + delta / 2
sigma = math.sqrt(2 * p_pool * (1 - p_pool))
n_base = math.ceil(((z_a + z_b) * sigma / delta) ** 2)
n_cuped = math.ceil(n_base * (1 - rho_cuped ** 2))
deff = 1 + (avg_cluster_size - 1) * icc
n_final = math.ceil(n_cuped * deff)
return {
"n_per_arm_unadjusted": n_base,
"n_per_arm_with_cuped": n_cuped,
"n_per_arm_final": n_final,
"n_clusters_needed": math.ceil(n_final / avg_cluster_size) if avg_cluster_size > 1 else None,
"design_effect": round(deff, 3),
"variance_reduction_pct": round(rho_cuped ** 2 * 100, 1),
}
# ── Deterministic assignment ──────────────────────────────────────────────────
def assign_variant(
unit_id: str,
experiment_id: str,
weights: dict[str, float] | None = None,
) -> str:
"""
Deterministic assignment via SHA-256 bucket keyed on experiment+unit.
Keying on both ensures the same user is not always in treatment across experiments.
Supports multi-arm splits via a weights dict that must sum to 1.0.
"""
weights = weights or {"control": 0.5, "treatment": 0.5}
assert abs(sum(weights.values()) - 1.0) < 1e-9, "Weights must sum to 1"
key = f"{experiment_id}:{unit_id}".encode()
bucket = int(hashlib.sha256(key).hexdigest(), 16) % 1_000_000 / 1_000_000
cumulative = 0.0
for arm, w in weights.items():
cumulative += w
if bucket < cumulative:
return arm
return list(weights)[-1] # floating-point safety valve
# ── Example ───────────────────────────────────────────────────────────────────
print(sample_size_service(0.45, 0.02, rho_cuped=0.72))
# {'n_per_arm_unadjusted': 19652, 'n_per_arm_with_cuped': 10991, ...}
print(assign_variant("user_8675309", "checkout_v2_2026q2")) # same every time
print(assign_variant("user_8675309", "onboarding_v3_2026q2")) # different experimentEnjoying these notes?
Get new lessons delivered to your inbox. No spam.