Decision Trees & Ensembles
Gradient boosted trees — not neural networks — dominate structured tabular data in production. XGBoost and LightGBM power fraud detection, insurance pricing, and ride-ETA prediction across the industry. Their edge over neural nets on tabular data comes from three properties: no normalization required (trees are scale-invariant), built-in feature importance (a single .feature_importances_ call), and competitive accuracy on datasets under 1M rows with minimal tuning. This lesson derives the impurity criteria, explains why bagging decorrelates trees to reduce variance while boosting reduces bias by fitting residuals, and walks through both methods on the Titanic dataset.
Theory
A decision tree is a sequence of yes/no questions that narrows down the answer. The diagram above shows the splits: each internal node asks a question about one feature, each branch is the answer, and each leaf is a prediction. The tree learns which questions to ask and in what order by finding the split that best separates the classes at each node. A single deep tree memorizes the training data; an ensemble of many shallow trees each trained on different data subsets averages out the noise.
Gini Impurity
For a node with class distribution :
A pure node has Gini = 0. A perfectly mixed binary node has Gini = 0.5. At each split, we pick feature and threshold to minimize the weighted Gini of the two children:
Entropy and Information Gain
An alternative split criterion based on Shannon entropy:
Both Gini and entropy give similar results in practice. Gini is faster to compute (no log).
Gini and entropy produce nearly identical splits in practice because both measure the same thing: purity. The key is that any split criterion must be concave in the class probabilities to guarantee that splitting always reduces impurity. A linear criterion would allow splits that don't actually improve separation. Concavity is the mathematical requirement that forces the U-shape of impurity measures — pure nodes score 0 and perfectly mixed nodes score maximum, with strict improvement guaranteed at every split.
Why Trees Overfit
An unconstrained tree grows until every leaf is pure — perfectly memorizing training data. With 30 features and 569 samples, an unlimited tree achieves 100% train accuracy but ~75–80% test accuracy. Limiting max_depth=4 gives ~94% test accuracy with much lower variance.
Bagging → Random Forest
Bootstrap aggregation: train trees, each on a bootstrap sample (sampled with replacement, ~63% unique rows). Average predictions:
Random forests add feature subsampling: each split considers only of features. This decorrelates the trees — averaging correlated predictors barely reduces variance, but decorrelated ones do.
Variance of the mean of i.i.d. estimators with variance : . Trees aren't i.i.d. (same data), but decorrelation via feature subsampling gets us most of this variance reduction.
Gradient Boosting
Build trees sequentially, each fitting the negative gradient of the loss:
where minimizes:
For MSE loss, the negative gradient equals the residuals . For log loss, it fits pseudo-residuals — the gradient of the loss with respect to the current prediction.
Walkthrough
Dataset: Kaggle Titanic (891 samples, binary survival prediction)
Feature Engineering
import pandas as pd
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.model_selection import cross_val_score
import numpy as np
df = pd.read_csv('train.csv')
df['FamilySize'] = df['SibSp'] + df['Parch'] + 1
df['IsAlone'] = (df['FamilySize'] == 1).astype(int)
df['Title'] = df['Name'].str.extract(r' ([A-Za-z]+)\.', expand=False)
df['Title'] = df['Title'].replace(
['Lady', 'Countess', 'Capt', 'Col', 'Don', 'Dr', 'Major', 'Rev', 'Sir', 'Jonkheer'],
'Rare'
)
df['Age'] = df.groupby(['Sex', 'Pclass'])['Age'].transform(
lambda x: x.fillna(x.median())
)
df['Sex'] = (df['Sex'] == 'male').astype(int)
features = ['Pclass', 'Sex', 'Age', 'Fare', 'FamilySize', 'IsAlone']
X = df[features]
y = df['Survived']Model Comparison
rf = RandomForestClassifier(n_estimators=200, max_depth=6, min_samples_leaf=4, random_state=42)
gb = GradientBoostingClassifier(n_estimators=200, max_depth=4, learning_rate=0.05, random_state=42)
rf_cv = cross_val_score(rf, X, y, cv=5, scoring='roc_auc')
gb_cv = cross_val_score(gb, X, y, cv=5, scoring='roc_auc')
print(f"RF AUC: {rf_cv.mean():.4f} ± {rf_cv.std():.4f}") # 0.8612 ± 0.0201
print(f"GB AUC: {gb_cv.mean():.4f} ± {gb_cv.std():.4f}") # 0.8789 ± 0.0185Gradient boosting wins — it explicitly fits residuals rather than averaging independent trees.
Code Implementation
train.pyfrom sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import roc_auc_score, classification_report
import joblib, numpy as np
def train(X, y, n_estimators=200, max_depth=8):
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42, stratify=y
)
model = RandomForestClassifier(
n_estimators=n_estimators,
max_depth=max_depth,
min_samples_leaf=4,
max_features='sqrt',
n_jobs=-1,
random_state=42,
)
model.fit(X_train, y_train)
y_prob = model.predict_proba(X_test)[:, 1]
auc = roc_auc_score(y_test, y_prob)
print(f"Test AUC: {auc:.4f}")
joblib.dump(model, "artifacts/rf_model.pkl")
return model
# Feature importance
importances = model.feature_importances_
# Fare: 0.31, Age: 0.27, Pclass: 0.22, FamilySize: 0.14Analysis & Evaluation
Where Your Intuition Breaks
More trees in a random forest always improve performance — variance keeps decreasing with ensemble size. This is true for variance, but bagging cannot reduce bias. If your base trees are systematically wrong about a region of the feature space (high bias), averaging hundreds of them produces a confident wrong answer, not a correct one. Adding trees helps until the variance is negligible; after that, the irreducible bias floor dominates and accuracy plateaus. The plateau is a signal that the base learner is too weak, not that you need more trees.
Hyperparameter Sensitivity
| Parameter | Low value | High value | Sweet spot |
|---|---|---|---|
n_estimators | High variance | Diminishing returns | 100–500 |
max_depth | Underfitting | Overfitting | 4–10 |
min_samples_leaf | Overfitting | Underfitting | 2–10 |
max_features | High bias | High variance | sqrt(p) |
Feature importance reveals which signals drive the ensemble's decisions. On the Titanic dataset, ticket fare and age dominate — consistent with the historical record that wealthier passengers and children had better access to lifeboats.
import matplotlib.pyplot as plt
import numpy as np
rf.fit(X, y) # refit on full training data
feature_names = ['Pclass', 'Sex', 'Age', 'Fare', 'FamilySize', 'IsAlone']
importances = rf.feature_importances_
idx = np.argsort(importances)
plt.figure(figsize=(6, 4))
plt.barh(np.array(feature_names)[idx], importances[idx], color='#6366f1')
plt.xlabel('Mean Decrease in Impurity')
plt.title('Random Forest Feature Importance — Titanic')
plt.tight_layout(); plt.savefig('feature_importance.png', dpi=150)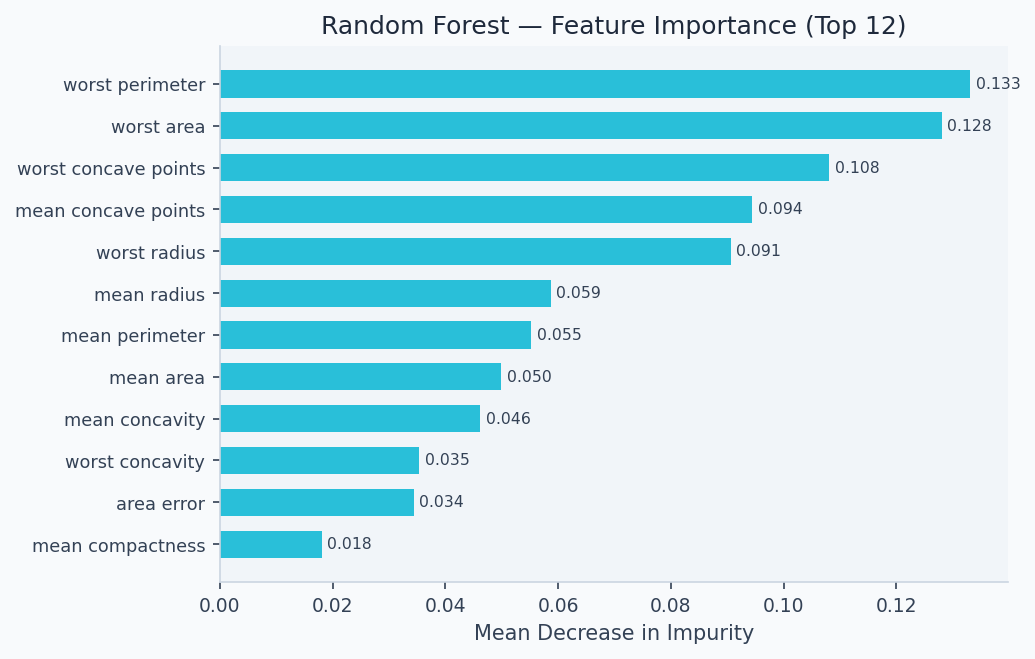
sklearn's mean-decrease-impurity importance is biased toward high-cardinality features. For reliable rankings, use permutation importance (sklearn.inspection.permutation_importance) or SHapley Additive exPlanations (SHAP) values.
Early Stopping in Gradient Boosting
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split
gb = GradientBoostingClassifier(
n_estimators=1000,
learning_rate=0.05,
subsample=0.8, # stochastic GB — reduces variance
validation_fraction=0.1,
n_iter_no_change=20, # early stopping patience
random_state=42,
)
gb.fit(X_train, y_train)
print(f"Stopped at {gb.n_estimators_} trees") # e.g., 347Production-Ready Code
serve_api/app.pyfrom fastapi import FastAPI
from pydantic import BaseModel
import joblib, numpy as np
app = FastAPI(title="Random Forest API")
model = joblib.load("artifacts/rf_model.pkl")
class TitanicFeatures(BaseModel):
Pclass: int
Sex: int # 1=male, 0=female
Age: float
Fare: float
FamilySize: int
IsAlone: int
@app.post("/predict")
def predict(req: TitanicFeatures):
x = np.array([[req.Pclass, req.Sex, req.Age, req.Fare, req.FamilySize, req.IsAlone]])
prob = float(model.predict_proba(x)[0][1])
return {
"survived_probability": round(prob, 4),
"prediction": int(prob > 0.5),
"confidence": "high" if abs(prob - 0.5) > 0.3 else "low",
}
@app.get("/health")
def health():
return {"status": "ok"}Track input feature distributions (PSI — population stability index) and prediction distribution over time. A PSI > 0.2 signals significant drift requiring retraining. Log feature values at inference time to enable drift detection.
Enjoying these notes?
Get new lessons delivered to your inbox. No spam.