Propensity Scores & Modern Methods
When you have rich observational data and believe you've measured all relevant confounders, propensity scores collapse the high-dimensional matching problem into a single scalar: the probability a unit would have been treated given its covariates. Weighting by inverse propensity recreates the equivalent of a randomized experiment from observational data — but only if the weights are well-behaved. Double ML extends this to the case where hundreds of features need to be controlled for, using cross-fitting to avoid regularization bias on the treatment effect estimate. Synthetic control handles the case where the treatment is a single unit (a city, a country, a product line) by constructing a weighted combination of control units that best matches its pre-treatment history. Each method handles a different regime of the observational inference problem, and each fails in a different way when its assumptions are violated — which is why sensitivity analysis (Rosenbaum bounds) is not optional.
Theory
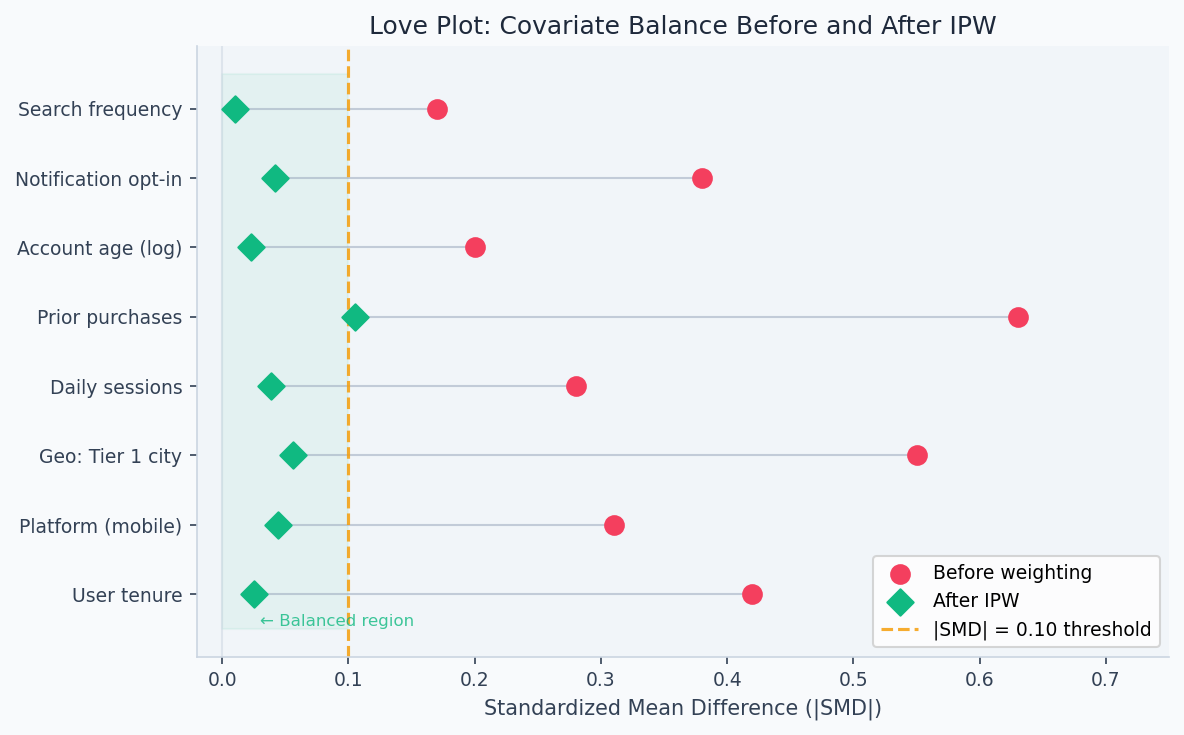
Each row is one covariate. After propensity score weighting, all SMDs fall inside ±0.1 (dashed lines) — treated and control groups are now comparable on observables.
In an observational study, treated and control units differ not just in treatment but in everything that made them more or less likely to be treated. Older patients get more aggressive treatment; wealthier customers get premium products. The propensity score compresses all those confounders into a single number — the probability of treatment — making it possible to compare units with the same probability of being treated but different actual assignments.
Propensity scores
When you have rich observational data and believe unconfoundedness holds — all confounders are measured — propensity scores provide a dimensionality reduction for causal estimation.
The propensity score is:
The propensity score works as a sufficient statistic for the full covariate vector because the balancing property is a consequence of probability theory, not an additional assumption. If two units have the same , their covariate distributions are identical in expectation — so any remaining difference in outcomes between treated and control units at that propensity score level must reflect the treatment effect, not selection bias. This is what makes a scalar summary of all confounders sufficient for identification, even when the covariate space is high-dimensional.
Balancing score theorem (Rosenbaum and Rubin, 1983): conditioning on balances the full covariate vector between treated and control groups:
Combined with unconfoundedness , this gives:
You only need to condition on a scalar rather than the full covariate vector — a dramatic simplification, especially in high dimensions.
Overlap assumption: for propensity score methods to work, every unit must have a positive probability of both treatment and control:
Violations create extreme weights (), inflating variance. Trim units with or (common: ). This buys variance reduction at the cost of external validity (you're excluding non-comparable units).
Inverse Probability Weighting (IPW)
IPW reweights the observed sample to make it representative of the target population. The Horvitz-Thompson estimator:
For treated units: weight — rare treated units (low ) get high weight, making them represent the full population. For control units: weight .
IPW is consistent when the propensity score model is correct. But it is sensitive to model misspecification — a slightly wrong model produces extreme weights and high-variance estimates.
After fitting IPW weights, the standard diagnostic is a Love plot — standardized mean differences (SMD) for each covariate before and after weighting. Values below |SMD| = 0.10 indicate good balance. The figure below shows what successful IPW looks like: pre-weighting SMDs as high as 0.63 on "prior purchases" collapse to near zero post-weighting across all features.
Doubly robust estimation
The doubly robust (DR) estimator combines IPW with outcome regression. It is consistent if either the propensity score model or the outcome model is correctly specified — not necessarily both.
When the outcome models are correct, the IPW residual terms have expectation zero — the estimator is just the outcome regression part. When the propensity score is correct, the IPW residuals correct for any outcome model misspecification. The two models "doubly protect" each other.
Double Machine Learning
Double Machine Learning (Double ML) (Chernozhukov et al., 2018) handles high-dimensional confounders via Neyman orthogonality. The idea: residualize both and on using flexible ML, then regress the residuals on each other.
Step 1 (cross-fitting): on held-out folds, estimate:
Step 2: regress on :
The Neyman orthogonality condition ensures that nuisance estimation errors enter the score only at second order — so is -consistent even when and converge at slower rates (e.g., ). This is the key advantage over naive plug-in estimation.
Synthetic Control
When there is only one treated unit (a country, a product line, a city) and several potential controls, synthetic control constructs a weighted combination of controls that matches the treated unit's pre-treatment trajectory:
Weights are chosen to minimize pre-treatment fit. Post-treatment, the synthetic control is the counterfactual. The treatment effect is the gap between the treated unit and its synthetic version.
Inference via placebo tests: with one treated unit, classical standard errors don't apply. Apply the same synthetic control procedure to each donor unit (as if it were treated). The p-value is the fraction of donors whose post/pre MSPE ratio exceeds the treated unit's.
Rosenbaum sensitivity analysis
After finding a significant effect from an observational study, how robust is it to unobserved confounding? Rosenbaum bounds quantify this.
Allow two matched units with identical observed covariates to differ in treatment odds by at most :
At : no hidden bias. For each , compute the worst-case p-value. The sensitivity threshold is the smallest that overturns your conclusion.
means: a confounder that multiplied treatment odds by 1.5 would make the result insignificant. means: you'd need a confounder tripling the treatment odds to explain away the effect. Large → robust result.
Walkthrough
Doubly robust ATE with cross-fitting
import numpy as np
from sklearn.ensemble import GradientBoostingClassifier, GradientBoostingRegressor
from sklearn.model_selection import KFold, cross_val_predict
from scipy import stats
def doubly_robust_ate(
X: np.ndarray,
T: np.ndarray,
Y: np.ndarray,
n_folds: int = 5,
) -> dict:
"""
Doubly robust ATE with cross-fitting.
Cross-fitting prevents nuisance model overfitting from biasing the ATE estimate.
"""
n = len(Y)
# Cross-fit propensity score e(X) = P(T=1 | X)
ps_model = GradientBoostingClassifier(n_estimators=100, max_depth=3, random_state=42)
e_hat = cross_val_predict(ps_model, X, T, cv=n_folds, method='predict_proba')[:, 1]
e_hat = np.clip(e_hat, 0.05, 0.95) # Trim extreme weights
# Cross-fit outcome models mu_1(X) = E[Y|T=1,X] and mu_0(X) = E[Y|T=0,X]
mu1_hat = np.zeros(n)
mu0_hat = np.zeros(n)
kf = KFold(n_splits=n_folds, shuffle=True, random_state=42)
for train_idx, val_idx in kf.split(X):
X_tr, T_tr, Y_tr = X[train_idx], T[train_idx], Y[train_idx]
m1 = GradientBoostingRegressor(n_estimators=100, random_state=42)
m0 = GradientBoostingRegressor(n_estimators=100, random_state=42)
m1.fit(X_tr[T_tr == 1], Y_tr[T_tr == 1])
m0.fit(X_tr[T_tr == 0], Y_tr[T_tr == 0])
mu1_hat[val_idx] = m1.predict(X[val_idx])
mu0_hat[val_idx] = m0.predict(X[val_idx])
# Doubly robust scores
dr_scores = (
T * (Y - mu1_hat) / e_hat
- (1 - T) * (Y - mu0_hat) / (1 - e_hat)
+ mu1_hat - mu0_hat
)
tau = dr_scores.mean()
se = dr_scores.std() / np.sqrt(n)
return {
"ate": round(tau, 4),
"se": round(se, 4),
"ci_95": (round(tau - 1.96*se, 4), round(tau + 1.96*se, 4)),
"p_value": round(2 * (1 - stats.norm.cdf(abs(tau/se))), 5),
}Propensity score diagnostics
Before interpreting any propensity score estimate, verify that the model achieves covariate balance. The standardized mean difference (SMD) should be below 0.1 for all covariates after weighting:
import pandas as pd
import numpy as np
def compute_smd(
df: pd.DataFrame,
covariates: list,
treatment_col: str,
weights: np.ndarray = None,
) -> pd.DataFrame:
"""Standardized mean differences before/after IPW weighting."""
treated = df[treatment_col] == 1
control = df[treatment_col] == 0
results = []
for cov in covariates:
x_t = df.loc[treated, cov].values
x_c = df.loc[control, cov].values
pooled_sd = np.sqrt((x_t.var() + x_c.var()) / 2 + 1e-10)
smd_raw = (x_t.mean() - x_c.mean()) / pooled_sd
smd_w = None
if weights is not None:
w_t, w_c = weights[treated.values], weights[control.values]
smd_w = (np.average(x_t, weights=w_t) - np.average(x_c, weights=w_c)) / pooled_sd
results.append({
'covariate': cov,
'smd_raw': round(smd_raw, 3),
'smd_weighted': round(smd_w, 3) if smd_w is not None else None,
'balanced': abs(smd_w or smd_raw) < 0.1,
})
return pd.DataFrame(results).sort_values('smd_raw', key=abs, ascending=False)
# Compute IPW weights and check balance
from sklearn.linear_model import LogisticRegression
ps = LogisticRegression(max_iter=500).fit(X, T)
e_hat = ps.predict_proba(X)[:, 1]
ipw_weights = np.where(T == 1, 1/e_hat, 1/(1-e_hat))
smd = compute_smd(df, covariates, 'treated', weights=ipw_weights)
print(smd)
# Aim for all |smd_weighted| < 0.1 before trusting the IPW estimateSynthetic control with placebo inference
from scipy.optimize import minimize
import numpy as np
def synthetic_control(
Y_treated: np.ndarray,
Y_donors: np.ndarray,
T0: int,
) -> dict:
"""
Synthetic control: find donor weights minimizing pre-treatment fit.
T0: number of pre-treatment periods.
"""
y_pre, X_pre = Y_treated[:T0], Y_donors[:T0]
J = X_pre.shape[1]
result = minimize(
lambda w: np.sum((y_pre - X_pre @ w)**2),
x0=np.ones(J)/J,
bounds=[(0, 1)]*J,
constraints={'type': 'eq', 'fun': lambda w: w.sum() - 1},
)
w = result.x
y_synth = Y_donors @ w
effects = Y_treated[T0:] - y_synth[T0:]
return {
"weights": w,
"y_synthetic": y_synth,
"att": effects.mean(),
"pre_fit_rmse": float(np.sqrt(np.mean((y_pre - y_synth[:T0])**2))),
"post_effects": effects,
}
def placebo_p_value(
Y_treated: np.ndarray,
Y_donors: np.ndarray,
T0: int,
) -> float:
"""
Placebo inference: apply synthetic control to each donor unit.
P-value = fraction of donors with larger post/pre MSPE ratio than the treated unit.
"""
def mspe_ratio(y_unit, y_other, T0):
r = synthetic_control(y_unit, y_other, T0)
pre_mspe = np.mean((y_unit[:T0] - r['y_synthetic'][:T0])**2)
post_mspe = np.mean((y_unit[T0:] - r['y_synthetic'][T0:])**2)
return post_mspe / (pre_mspe + 1e-10)
J = Y_donors.shape[1]
treated_ratio = mspe_ratio(Y_treated, Y_donors, T0)
# Placebo: treat each donor as if it were the treated unit
placebo_ratios = []
for j in range(J):
other_donors = np.delete(Y_donors, j, axis=1)
r = mspe_ratio(Y_donors[:, j], other_donors, T0)
placebo_ratios.append(r)
p_val = np.mean(np.array(placebo_ratios) >= treated_ratio)
return round(p_val, 3)Rosenbaum sensitivity bounds
from scipy import stats
def rosenbaum_sensitivity(
treated_outcomes: np.ndarray,
control_outcomes: np.ndarray,
gamma_range: list = None,
) -> pd.DataFrame:
"""
Sensitivity analysis for matched pairs under Rosenbaum's model.
gamma_range: list of Gamma values to test.
"""
if gamma_range is None:
gamma_range = [1.0, 1.25, 1.5, 2.0, 2.5, 3.0]
diffs = treated_outcomes - control_outcomes
n = len(diffs)
ranks = stats.rankdata(np.abs(diffs))
T_obs = sum(r for r, d in zip(ranks, diffs) if d > 0)
rows = []
for gamma in gamma_range:
p_plus = gamma / (1 + gamma)
E_T = n*(n+1)/2 * p_plus
Var_T = n*(n+1)*(2*n+1)/6 * p_plus*(1-p_plus)
z = (T_obs - E_T) / np.sqrt(Var_T)
p_worst = 1 - stats.norm.cdf(z)
rows.append({
'gamma': gamma,
'p_worst_case': round(p_worst, 5),
'conclusion_holds': p_worst < 0.05,
})
return pd.DataFrame(rows)
# Interpret: find gamma* — the smallest Gamma where conclusion_holds = False
# gamma*=1.5 → fragile; gamma*=3.0 → robust to large hidden confoundersAnalysis & Evaluation
Where Your Intuition Breaks
A good Love plot (balanced covariates after weighting) means the causal estimate is valid. Covariate balance is a necessary condition for unconfoundedness, not a sufficient one. A Love plot only shows balance on measured covariates — unmeasured confounders remain unbalanced regardless of how well the propensity model performs on observed features. Rosenbaum sensitivity analysis quantifies how large an unmeasured confounder would need to be to reverse the conclusion, but it cannot rule out confounding. Perfect balance on observables with a hidden confounder still produces a biased estimate.
Method selection guide
| Situation | Method | Identifying assumption |
|---|---|---|
| Random assignment possible | RCT | None |
| Phased rollout / regional variation | DiD | Parallel trends |
| Threshold-based eligibility | RDD | Continuity at cutoff |
| Valid instrument available | IV / 2SLS | Exclusion restriction |
| Single treated unit | Synthetic Control | Pre-treatment fit |
| Rich covariates, unconfoundedness plausible | DR / Double ML | No unmeasured confounders |
When multiple methods are available, triangulate. If DiD, IV, and doubly robust all give similar estimates, that's much stronger evidence than any single method alone.
Assumption severity
Not all identifying assumptions are equally testable or plausible:
| Assumption | Testable? | Severity if violated |
|---|---|---|
| Parallel trends (DiD) | Partially (pre-period) | Moderate — can add unit trends |
| Continuity (RDD) | Yes (McCrary, covariate continuity) | Severe — invalidates RDD entirely |
| Exclusion restriction (IV) | No — requires economic reasoning | Severe — biases 2SLS toward OLS |
| No unmeasured confounders | No — requires domain knowledge | Severe — quantify with Rosenbaum bounds |
| Overlap (PS methods) | Yes — check propensity score distribution | Moderate — trim and note external validity loss |
Causal inference from observational data is not as reliable as a randomized experiment. Observational estimates carry implicit assumptions that cannot be verified from data alone. When you report an observational effect, always state which assumption you're relying on, how you've tried to validate it, and what a Rosenbaum sensitivity analysis says about robustness.
End-to-end observational pipeline
from dataclasses import dataclass, field
@dataclass
class CausalResult:
method: str
ate: float
ci_lower: float
ci_upper: float
p_value: float
n_units: int
validated: dict = field(default_factory=dict)
warnings: list = field(default_factory=list)
def observational_pipeline(df, outcome, treatment, covariates):
"""Standard pipeline: overlap check → balance check → DR estimation → validation."""
warnings, validated = [], {}
X, T, Y = df[covariates].values, df[treatment].values, df[outcome].values
# 1. Fit propensity score and check overlap
from sklearn.linear_model import LogisticRegression
ps = LogisticRegression(max_iter=500).fit(X, T)
e_hat = ps.predict_proba(X)[:, 1]
poor_overlap = ((e_hat < 0.05) | (e_hat > 0.95)).mean()
if poor_overlap > 0.05:
warnings.append(f"Overlap: {poor_overlap:.1%} of units outside [0.05, 0.95]")
validated['overlap'] = poor_overlap < 0.05
# 2. Pre-treatment balance
smd = compute_smd(df, covariates, treatment)
max_smd = smd['smd_raw'].abs().max()
if max_smd > 0.25:
warnings.append(f"High pre-treatment imbalance: max SMD = {max_smd:.2f}")
# 3. Doubly robust ATE
res = doubly_robust_ate(X, T, Y)
# 4. Post-weighting balance check
ipw_w = np.where(T == 1, 1/e_hat, 1/(1-e_hat))
smd_after = compute_smd(df, covariates, treatment, weights=ipw_w)
max_smd_after = smd_after['smd_weighted'].abs().max()
validated['balance'] = max_smd_after < 0.1
if max_smd_after >= 0.1:
warnings.append(f"Post-weighting imbalance: max SMD = {max_smd_after:.2f}")
return CausalResult(
method="doubly_robust",
ate=res['ate'], ci_lower=res['ci_95'][0], ci_upper=res['ci_95'][1],
p_value=res['p_value'], n_units=len(df),
validated=validated, warnings=warnings,
)In platforms where A/B tests can't be run (supply-side features, city-level policy changes), causal inference from observational data is the only option. Best practice: run at least two independent methods (e.g., DiD and synthetic control), plot event study diagnostics to validate parallel trends, and report Rosenbaum bounds alongside the estimate. An observational result that hasn't been validated with multiple methods should be treated as exploratory and not used for product decisions.
Production-Ready Code
Doubly-robust AIPW is the production-grade default when you have enough covariates to worry about model misspecification. The key operational concern is weight diagnostics: extreme IPW weights signal overlap failures that inflate variance. Cross-fitting is mandatory — in-sample nuisance estimation introduces regularisation bias that contaminates the treatment effect estimate.
# production_propensity.py
# AIPW (doubly-robust) ATE estimation with weight diagnostics and overlap trimming.
from __future__ import annotations
import math
import numpy as np
import scipy.stats as stats
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.model_selection import KFold
from sklearn.preprocessing import StandardScaler
def weight_diagnostics(e_hat: np.ndarray, T: np.ndarray) -> dict:
"""
Run BEFORE any weighted estimator.
Extreme weights (max > 20 or ESS < 30% of n) signal overlap failure.
"""
w = np.where(T == 1, 1.0 / e_hat, 1.0 / (1.0 - e_hat))
ess = w.sum() ** 2 / (w ** 2).sum()
return {
"max_weight": round(float(w.max()), 2),
"p99_weight": round(float(np.percentile(w, 99)), 2),
"effective_sample_size": round(float(ess), 1),
"ess_fraction": round(float(ess / len(T)), 3),
"min_propensity": round(float(e_hat.min()), 4),
"max_propensity": round(float(e_hat.max()), 4),
"overlap_warning": bool(e_hat.min() < 0.05 or e_hat.max() > 0.95),
"action": (
"Trim units with e < 0.05 or e > 0.95 before weighting"
if e_hat.min() < 0.05 or e_hat.max() > 0.95
else "OK"
),
}
def aipw_ate(
X: np.ndarray,
T: np.ndarray,
Y: np.ndarray,
n_folds: int = 5,
trim: float = 0.05,
) -> dict:
"""
Augmented IPW (doubly-robust) ATE via cross-fitting.
Doubly robust: consistent if EITHER the propensity OR outcome model is correct.
Cross-fitting removes regularisation bias: nuisance models are trained on
held-out folds so their errors are orthogonal to the treatment effect estimate.
"""
n = len(Y)
scaler = StandardScaler()
Xs = scaler.fit_transform(X)
e_hat = np.zeros(n)
mu1_hat = np.zeros(n)
mu0_hat = np.zeros(n)
kf = KFold(n_splits=n_folds, shuffle=True, random_state=42)
for train_idx, val_idx in kf.split(Xs):
X_tr, X_val = Xs[train_idx], Xs[val_idx]
T_tr = T[train_idx]
Y_tr = Y[train_idx]
ps = LogisticRegression(C=1.0, max_iter=500, random_state=42)
ps.fit(X_tr, T_tr)
e_hat[val_idx] = ps.predict_proba(X_val)[:, 1]
m1 = GradientBoostingRegressor(n_estimators=100, random_state=42)
m0 = GradientBoostingRegressor(n_estimators=100, random_state=42)
if T_tr.sum() >= 10:
m1.fit(X_tr[T_tr == 1], Y_tr[T_tr == 1])
mu1_hat[val_idx] = m1.predict(X_val)
else:
mu1_hat[val_idx] = Y_tr[T_tr == 1].mean()
if (1 - T_tr).sum() >= 10:
m0.fit(X_tr[T_tr == 0], Y_tr[T_tr == 0])
mu0_hat[val_idx] = m0.predict(X_val)
else:
mu0_hat[val_idx] = Y_tr[T_tr == 0].mean()
keep = (e_hat >= trim) & (e_hat <= 1 - trim)
e_k, mu1_k, mu0_k = e_hat[keep], mu1_hat[keep], mu0_hat[keep]
T_k, Y_k = T[keep], Y[keep]
psi = (
T_k * (Y_k - mu1_k) / e_k
- (1 - T_k) * (Y_k - mu0_k) / (1 - e_k)
+ mu1_k - mu0_k
)
tau = float(psi.mean())
se = float(psi.std() / math.sqrt(len(psi)))
z = tau / se
p_val = float(2 * (1 - stats.norm.cdf(abs(z))))
ci = (tau - 1.96 * se, tau + 1.96 * se)
return {
"ate_aipw": round(tau, 5),
"se": round(se, 5),
"ci_95": (round(ci[0], 5), round(ci[1], 5)),
"p_value": round(p_val, 6),
"n_trimmed": int((~keep).sum()),
"weight_diagnostics": weight_diagnostics(e_hat[keep], T_k),
}
# ── Example ───────────────────────────────────────────────────────────────────
rng = np.random.default_rng(42)
n = 2_000
X = rng.normal(0, 1, (n, 5))
propensity = 1 / (1 + np.exp(-X[:, 0]))
T = rng.binomial(1, propensity).astype(float)
Y = 2.5 * T + X[:, 0] + rng.normal(0, 1, n) # true ATE = 2.5
result = aipw_ate(X, T, Y)
print(result)
# ate_aipw ≈ 2.5, overlap_warning: FalseEnjoying these notes?
Get new lessons delivered to your inbox. No spam.